-
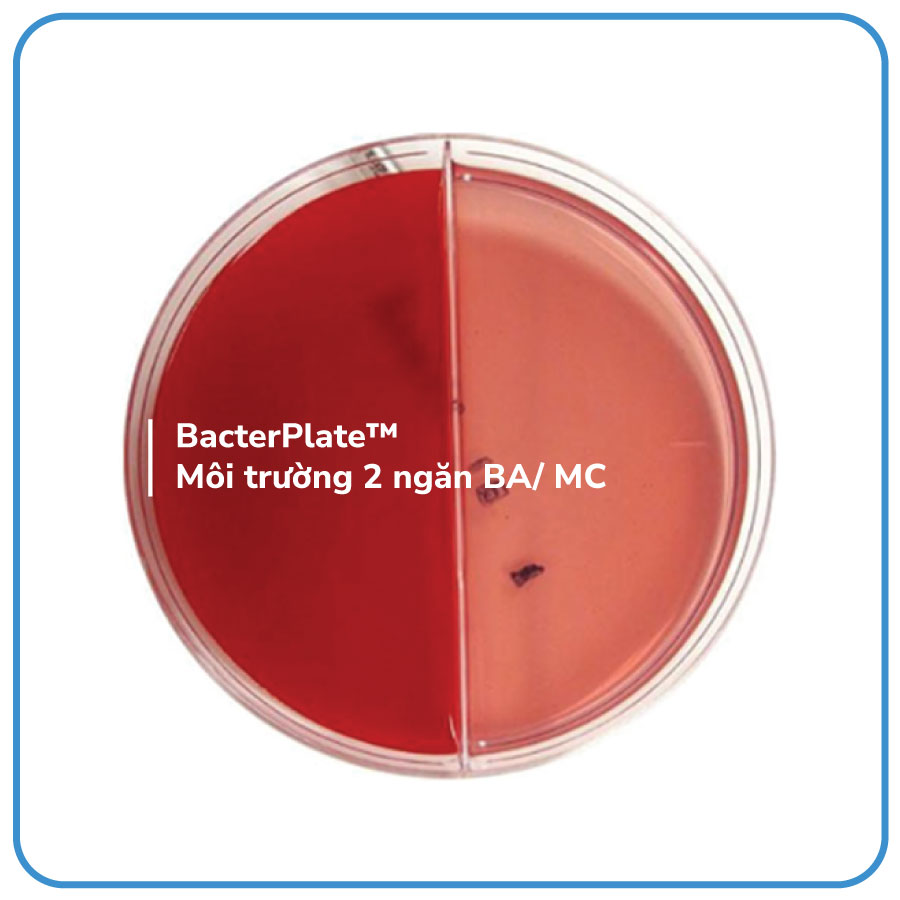
TraceMedia™ Columbia Sheep/Macconkey (BA/MC 90mm) is a dual-compartment culture medium used for the cultivation and isolation of pathogenic bacteria from clinical specimens.
PURPOSE
- TraceMedia™ Columbia Sheep/Macconkey (BA/MC 90mm) consists of two nutrient-rich compartments: Blood Agar (BA) and MacConkey (MC).
- BA: This medium is used for the isolation of most types of microorganisms. It also helps in differentiating the hemolytic types (alpha, beta, gamma).
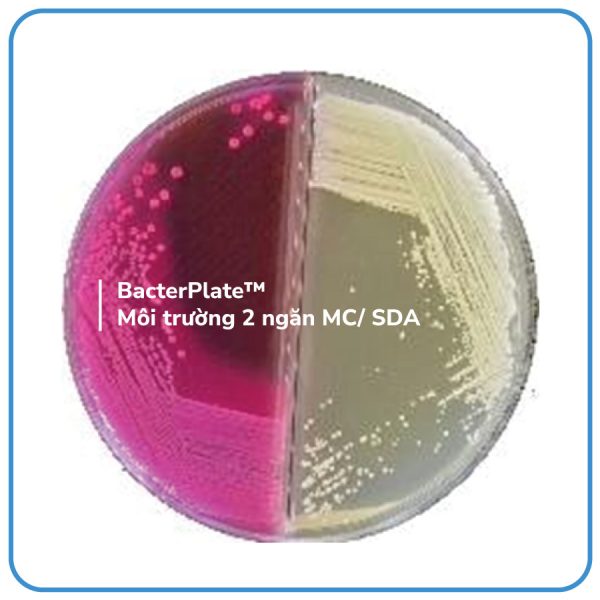
- MC: MacConkey agar is a selective medium used for the isolation of Gram-negative enteric bacteria that are easy to culture. It also helps in differentiating the lactose fermenters.
PRINCIPLE
TraceMedia™ Columbia Sheep/Macconkey (BA/MC 90mm)
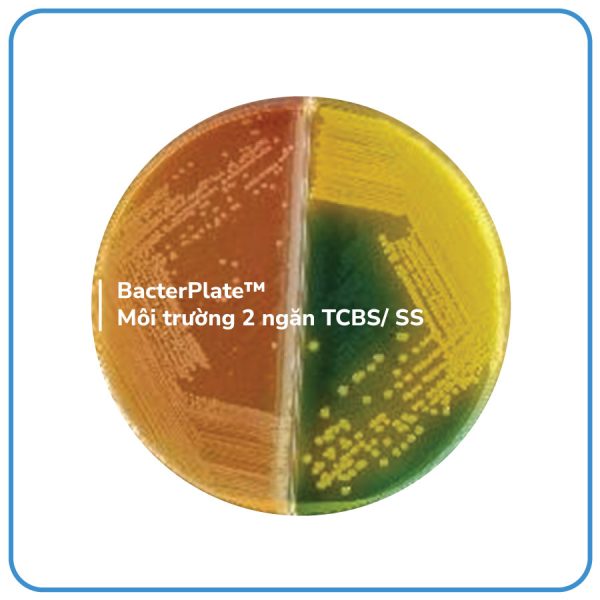
- Blood Agar (BA) compartment: The Columbia Agar Base supplemented with sheep blood provides a growth medium and allows for the determination of bacterial hemolytic reactions. On the surface of the blood agar plate, bacterial colonies exhibit different types of hemolysis. Beta hemolysis results in a clear zone of complete hemolysis, alpha hemolysis shows a greenish zone around the colonies, and gamma hemolysis shows no hemolysis.
- MacConkey (MC) compartment: MacConkey agar is used to selectively grow Gram-negative bacteria, particularly those belonging to the Enterobacteriaceae family and Pseudomonas genus. This medium inhibits the growth of Gram-positive bacteria and some difficult-to-culture Gram-negative bacteria due to the presence of crystal violet and bile salts in the medium. Lactose-fermenting bacteria produce acid, causing a decrease in the pH of the medium. As the pH decreases, the color of the medium changes to pink or red. Additionally, due to the acid produced from lactose fermentation reacting with bile salts, a pink halo is formed around the bacterial colonies. Lactose-weak fermenters also change the medium to pink or red, but no halo is formed around the colonies. Bacteria that do not ferment lactose will retain their original color on the medium, and the colonies will also be colorless.
READING RESULTS
- After incubation for the required time, typically 16-24 hours, observe the growth of bacterial colonies on the surface of the agar plate.
- To identify the isolated bacteria, further tests must be conducted using appropriate methods.


 Tiếng Việt
Tiếng Việt